
2nd MICCAI Workshop on
“Distributed And Collaborative Learning”
Important Dates
- Submission Deadline: 2nd of July 2021, 23:59 AOE
- Notification of Acceptance: 22th of July 2021
- Camera Ready: 30th of July 2021
- Workshop proceedings: 6th of August 2021
- Workshop Date: 1 October 2021, (virutal)
News
[June] Submission deadline was extended to 2nd of July due to several requests!
[June] NVIDIA sponsors a RTX 3090 for the best paper award!
[June] Our first keynote speaker is confirmed! We are happy to welcome a thought leader of federated learning: Peter Kairouz (Research Scientist, Google)
[June] MICCAI 2021 will take place as a virtual conference
[May] Submission is open on CMT system
[March] The DCL Workshop has been approved for MICCAI 2021
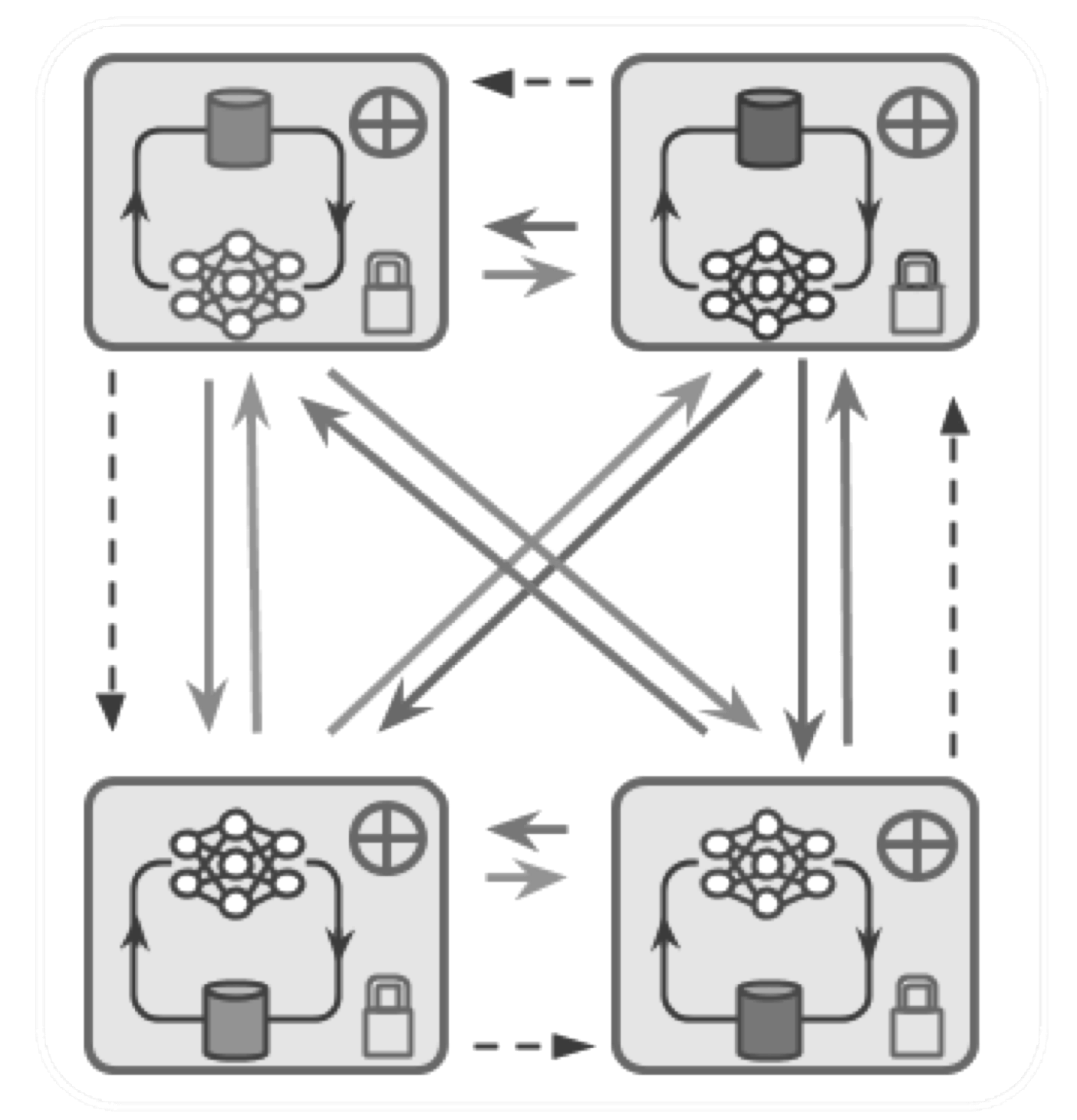
Distributed and Collaborative Learning
Deep learning empowers enormous scientific advances, with key applications in healthcare. It has been widely accepted that it is possible to achieve better models with growing amounts of data. However, enabling learning on these huge datasets or training huge models in a timely manner requires to distribute the learning on several devices. One particularity in the medical domain, and in the medical imaging setting is that data sharing across different institutions often becomes impractical due to strict privacy regulations, making the collection of large-scale centralized datasets practically impossible.
Some of the problems, therefore, become: how can we train models in a distributed way on several devices? And is it possible to achieve models as strong as those that can be trained on large centralized datasets without sharing data and breaching the restrictions on privacy and property? Distributed machine learning, including Federated Learning (FL) approaches, could be helpful to solve the latter problem. Different institutions can contribute to building more powerful models by performing collaborative training without sharing any training data. The trained model can be distributed across various institutions but not the actual data. We hope that with FL and other forms of distributed and collaborative learning, the objective of training better, more robust models of higher clinical utility while still protecting the privacy within the data can be achieved.
Call for Papers
Through the second MICCAI Workshop on Distributed And Collaborative Learning (DCL), we aim to provide a discussion forum to compare, evaluate and discuss methodological advancements and ideas around federated, distributed, and collaborative learning schemes that are applicable in the medical domain. Topics include but are not limited to:
- Federated, distributed learning, and other forms of collaborative learning
- Server-client and peer-to-peer learning
- Advanced data and model parallelism learning techniques
- Optimization methods for distributed or collaborative learning
- Privacy-preserving technique and security for distributed or collaborative learning
- Efficient communication and learning (multi-device, multi-node)
- Adversarial, inversion and other forms of attacks on distributed or collaborative learning
- Dealing with unbalanced (non-IID) data in collaborative learning
- Diverse decentralized medical imaging data analysis
- Security-auditing system for federated learning
- Asynchronous learning
- Software tools and implementations of distributed or collaborative learning
- Model sharing techniques, sparse/partial learning of models
- Applications of distributed/collaborative learning techniques: multi-task learning, model agnostic learning, meta-learning, etc.
Program
Keynote Session
Peter Kairouz
Google Research
Peter Kairouz is a research scientist at Google, where he leads research efforts on federated learning and privacy-preserving technologies. Before joining Google, he was a Postdoctoral Research Fellow at Stanford University. He received his Ph.D. in electrical and computer engineering from the University of Illinois at Urbana-Champaign (UIUC). He is the recipient of the 2012 Roberto Padovani Scholarship from Qualcomm's Research Center, the 2015 ACM SIGMETRICS Best Paper Award, the 2015 Qualcomm Innovation Fellowship Finalist Award, and the 2016 Harold L. Olesen Award for Excellence in Undergraduate Teaching from UIUC.
Topic: Privacy in Federated Learning at Scale
Abstract: I will start this talk by overviewing Federated Learning (FL) and its core data minimization principles. I will then describe how privacy can be strengthened and rigorized using complementary privacy techniques such as differential privacy, secure multi-party computation, and privacy auditing methods. I will spend much of the talk describing how we can carefully combine technologies like differential privacy and secure aggregation to obtain formal distributed privacy guarantees without fully trusting the server in adding noise. I will present a comprehensive end-to-end system, which appropriately discretizes the data and adds discrete Gaussian noise before performing secure aggregation. I will conclude by showing experimental results that demonstrate that our solution is able to achieve a comparable accuracy to central differential privacy (which requires trusting the server in adding noise) with just 16 bits of precision per value.

Marco Lorenzi
Inria
Marco Lorenzi is tenured research scientist at Inria Sophia Antipolis and Université Côte d’Azur, and holds a chair in the French Interdisciplinary Institutes of Artificial Intelligence (3IA). Prior to this, he was Research Associate at University College London. He got his PhD at Inria Sophia Antipolis working in the Asclepios Research Group. His research interests are in the development of statistical and machine learning methods for the analysis of large-scale and heterogeneous biomedical data. He currently leads Fed-BioMed, an initiative for the deployment of federated learning in clinical applications, funded by the French National Research Agency and the French National Artificial Intelligence Research Program.
Topic: Federated Learning in Healthcare: from Theory to Practice
Abstract: During this talk, I will present our research and development program for the deployment of federated learning (FL) in healthcare applications. From the theoretical perspective, I’ll propose novel statistical approaches to tackle the problem of data heterogeneity in FL, by investigating the theoretical properties of client sampling and aggregation mechanisms. Furthermore, I will introduce Bayesian paradigms for modelling clients heterogeneity in presence of data biases and missing information. From the practical perspective, I will illustrate the open source software Fed-BioMed, an Inria development initiative to enable scalable and general FL applications. I will present its basic FL paradigms and components, showcase its workflow for easily deploying models in typical FL scenarios, and illustrate real life examples of its deployment and use in multi-centric studies.

Meet the Organising Team
Contact: nrieke(at)nvidia.com
Shadi Albarqouni
Helmholtz AI & Technical University of Munich


M. Jorge Cardoso
King’s College London

Nicola Rieke
NVIDIA

Daguang Xu
NVIDIA
Spyridon Bakas
University of Pennsilvania

Bennett Landman
Vanderbilt University

Xiaoxiao Li
The University of British Columbia

Holger Roth
NVIDIA

Program Commitee
Aaron Carass, Johns Hopkins University
Amir Alansary, Johns Hopkins University
Andriy Myronenko, NVIDIA
Benjamin A Murray, King's College London
Christian Wachinger, LMU Munich
Daniel Rubin, Stanford Univesity
Dong Yang, NVIDIA Corporation
Ipek Oguz, Vanderbilt University
G Anthony Reina, Intel Corporation
Jayashree Kalpathy-Cramer, MGH/Harvard Medical School
Jonas Scherer, DKFZ
Jonny Hancox, NVIDIA
Kate Saenko, Boston University
Ken Chang, Massachusetts General Hospital
Khaled Younis, GE
Klaus Kades, DKFZ
Ling Shao, Inception Institute of Artificial Intelligence
JMarco Nolden, German Cancer Research Center
Maximilian Zenk, DKFZ
Meirui Jiang, CUHK
Micah J Sheller, Intel Corporation
Nir Neumark, Massachusetts General Hospital
Qiang Yang, HKUST
Quande Liu, CUHK
Quanzheng Li, Harvard Medical School/Massachusetts General Hospital
Ralf Floca, DKFZ
Reuben Dorent King's College London
Sarthak Pati, University of Pennsylvania
Shunxing Bao, Vanderbilt University
Walter Hugo Lopez Pinaya, KCL
Wojciech Samek, Fraunhofer HHI
Xingchao Peng, Boston University
Yang Liu, Webank
Yuankai Huo, Vanderbilt University
Zach Eaton-Rosen, King's College London
Zijun Huang, Columbia University
Ziyue Xu, NVIDIA
Aaron Carass, Johns Hopkins University
Amir Alansary, Johns Hopkins University
Andriy Myronenko, NVIDIA
Benjamin A Murray, King's College London
Christian Wachinger, LMU Munich
Daniel Rubin, Stanford Univesity
Dong Yang, NVIDIA Corporation
Ipek Oguz, Vanderbilt University
G Anthony Reina, Intel Corporation
Jayashree Kalpathy-Cramer, MGH/Harvard Medical School
Jonas Scherer, DKFZ
Jonny Hancox, NVIDIA
Kate Saenko, Boston University
Ken Chang, Massachusetts General Hospital
Khaled Younis, GE
Klaus Kades, DKFZ
Ling Shao, Inception Institute of Artificial Intelligence
JMarco Nolden, German Cancer Research Center
Maximilian Zenk, DKFZ
Meirui Jiang, CUHK
Micah J Sheller, Intel Corporation
Nir Neumark, Massachusetts General Hospital
Qiang Yang, HKUST
Quande Liu, CUHK
Quanzheng Li, Harvard Medical School/Massachusetts General Hospital
Ralf Floca, DKFZ
Reuben Dorent King's College London
Sarthak Pati, University of Pennsylvania
Shunxing Bao, Vanderbilt University
Walter Hugo Lopez Pinaya, KCL
Wojciech Samek, Fraunhofer HHI
Xingchao Peng, Boston University
Yang Liu, Webank
Yuankai Huo, Vanderbilt University
Zach Eaton-Rosen, King's College London
Zijun Huang, Columbia University
Ziyue Xu, NVIDIA

Sponsor
NVIDIA sponsors a RTX 3090 for the DCL best paper award!
Submission Guidelines
Format: Papers will be submitted electronically following Lecture Notes in Computer Science (LNCS) style of up to 8 + 2 pages (same as MICCAI 2021 format). Submissions exceeding page limit will be rejected without review. Latex style files can be found from Springer, which also contains Word instructions. The file format for submissions is Adobe Portable Document Format (PDF). Other formats will not be accepted.
Double Blind Review: DCL reviewing is double blind. Please review the Anonymity guidelines of MICCAI main conference, and confirm that the author field does not break anonymity.
Paper Submission: DCL uses the CMT system for online submission.
Supplemental Material: Supplemental material submission is optional, following same deadline as the main paper. Contents of the supplemental material would be referred to appropriately in the paper, while reviewers are not obliged to read them.
Submission Originality: Submissions should be original, no paper of substantially similar content should be under peer review or has been accepted for a publication elsewhere (conference/journal, not including archived work).
Proceedings: The proceedings of DCL 2021 will be published as part of the joint MICCAI Workshops proceedings with Springer (LNCS)

